What Is A Template In Pcr
The PCR (template) DNA must be a highly purified Deoxyribonucleic acid having 30ng to 50ng concentration, l% to 55% GC content and free from chemical contaminants and other DNA contaminants.
The PCR template DNA is ane of the of import ingredients for achieving a successful PCR reaction. It is as of import as the DNA primers, Taq DNA polymerase, dNTPs and PCR reaction buffer.
Inappropriate concentration fo the template DNA in PCR reaction results into reaction failure, likewise, a higher concentration of the DNA template in PCR, resulting in imitation positive results.
The Taq Dna polymerase tin only be activated if it recognises the template Deoxyribonucleic acid every bit a substrate for the PCR reaction. Although the polymerization cannot be started until the primer binds to the template Deoxyribonucleic acid.
Similarly, the dTNPs cannot be binds to the site of amplification until the Taq recognised the PCR template DNA as a substrate for the reaction.
Therefore, the DNA used equally a template in the PCR reaction plays an important part in distension.
In this article, we will discuss several properties of the PCR Dna along with its concentration required to perform the PCR reaction.
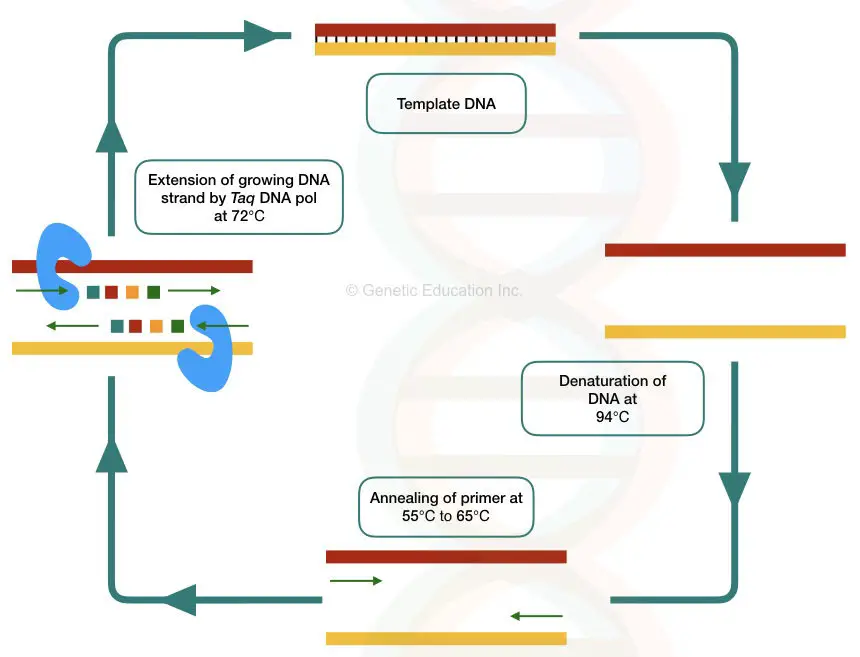
Introduction:
The template DNA used in the PCR is isolated using several different types of Dna extraction methods. These Deoxyribonucleic acid extraction methods are enlisted below,
- CTAB Deoxyribonucleic acid extraction method
- Phenol chloroform Deoxyribonucleic acid extraction.
- Proteinase Grand Deoxyribonucleic acid extraction method
Which types of Dna extraction method is selected for our protocol is depends on the types of tissue we are using,
For instance, the phenol-chloroform Dna extraction method is a good choice for extracting DNA from the blood. similarly, the CTAB DNA extraction method is the best selection for plant tissue.
After the Deoxyribonucleic acid extraction and earlier the polymerase concatenation reaction, the PCR (template) DNA must exist quantified considering the success of any PCR reaction depends on the quantity and the purity of the template DNA.
For quantification of the Deoxyribonucleic acid, we tin can use two methods spectrophotometric method and fluorometric method. However, the spectroscopic method can do both quantification and purity cheque.
The DNA used for the PCR tin be broadly divided into ii categories:
- Target sequence Dna (the amount of Dna to be amplified)
- Non- target sequence Deoxyribonucleic acid.
The target Deoxyribonucleic acid is generally a smaller portion of the template DNA which is selected by us for the distension, this may exist a short or long sequence or an entire gene.
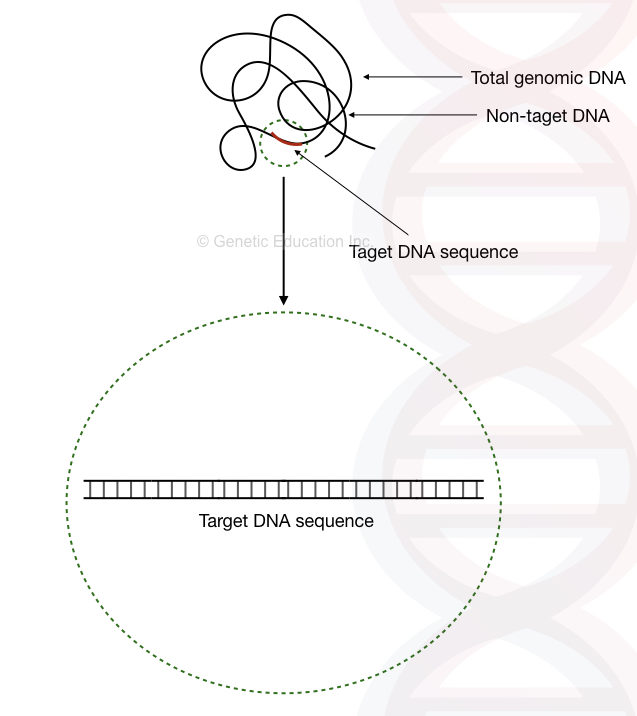
In general PCR reaction, the target Dna length is nearly 100bp to 1000bp or maximum 1500bp. For longer PCR which takes more fourth dimension and required advance reaction preparation, the target Deoxyribonucleic acid sequence may be 2000bp to 10,000bp long.
However, for the amplification of longer Dna fragment, more advanced protocol and other PCR additives are required.
The target sequence is < 1% of the full Deoxyribonucleic acid.
The balance of the DNA is non-target sequence DNA which cannot be amplified into the reaction or it is a kind of junk DNA.
All the same, the non-target Deoxyribonucleic acid can also hinder into the reaction.
Backdrop of PCR template DNA
The template Dna must be contaminant-free:
The genomic Dna used in the PCR reaction must exist contaminant free. Phenol, CTAB, other inorganic and organic salts are the common contaminant of the Dna.
Proper washing and Deoxyribonucleic acid purification are required before going to PCR.
If the purity of the Dna is above or below 1.8 (260/280 ratio), the PCR template DNA must exist contaminated with proteins, phenol or RNA.
In that example, purify Dna before doing PCR otherwise the PCR amplification may not be achieved.
The template DNA must be free from other Dna contaminants:
PCR template Dna contaminated with the DNA of other organisms results in the false positive amplification into the PCR reaction.
For example, if we are doing tuberculosis PCR, there is a adventure that along with the pathogen Deoxyribonucleic acid the genomic Dna of the host might be amplified.
In this case, the primers bind to the not-complementary or semi-complementary sequences of other DNA and amplify it.
False positive results are ane of the biggest bug in the PCR amplification. To avert information technology purify the DNA before PCR.
[wp_ad_camp_1]
GC content of the DNA:
The Dna is made upwards of the Adenine, Thymine, Cytosine and Guanine. Three hydrogen bonds between G and C and two hydrogen bonds between A and T joins two strands of Dna.
Due to more than hydrogen bonds between G and C, the amplification of GC rich Dna is difficult. To avoid this, select the region of target DNA with GC content between l% to 55%.
Higher GC content increases the melting temperature of the reaction, also information technology is very difficult to dilate it.
Concentration of template DNA:
30ng to 50ng Deoxyribonucleic acid is required for 25 microliter PCR reaction. if the concentration is lower, the DNA might not be amplified. However, amplification is reported even with the 10pg of Deoxyribonucleic acid.
Furthermore, a higher concentration of Dna also hinders in the PCR reaction. At that place are two possibilities for that, the Deoxyribonucleic acid may amplify (non-specific) or the Deoxyribonucleic acid may not exist amplified.
Due to the higher concentration of Dna, the PCR tube is filled with a college corporeality of non-specific DNA and hinders in the activity of the other reagents or the PCR reagents cannot be reached at the target Dna sequence which leads not-amplification and reaction failure.
In another possibility, due to the higher corporeality of the non-target template Deoxyribonucleic acid, the primer bind to other than its complementary sequences and results in fake positive PCR amplification.
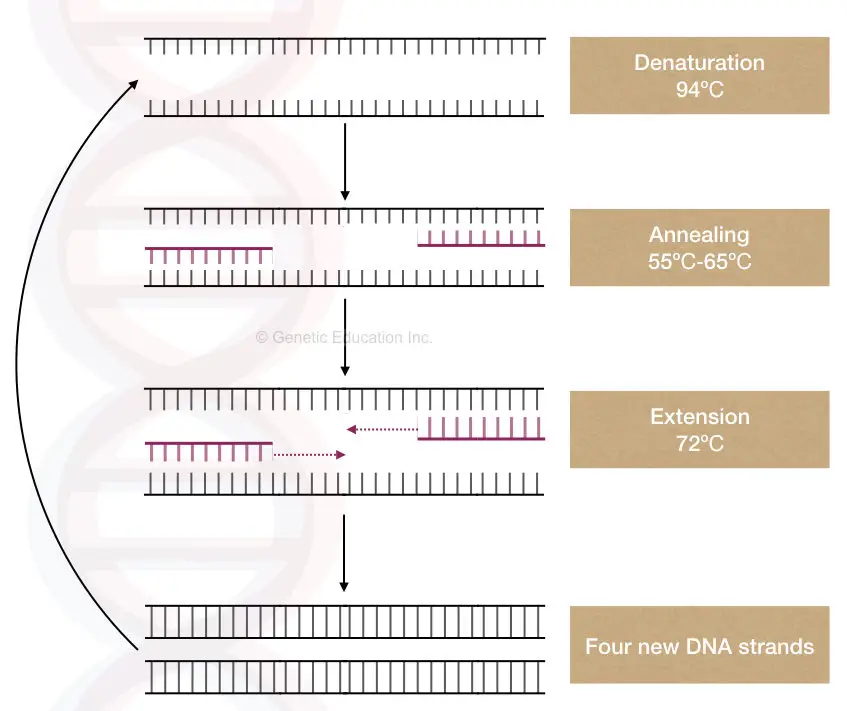
Purity of template DNA:
Another important property of PCR (template) Deoxyribonucleic acid is the purity of the DNA. The unpurified- contaminated Deoxyribonucleic acid cannot exist amplified.
Contaminants such as phenol, chloroform and other salt hinder in the PCR reaction. Phenol is one of common PCR inhibitor.
Tips: Ever dissolve DNA in TE buffer and the pH of the TE buffer must be nearly 8.0. The integrity of the Dna maintained at the pH ~8.0.
Also, bank check the templet DNA if it is fragmented or not, fragmented Dna cannot exist used in the PCR. For checking that use agarose gel electrophoresis technique.
If the concentration of the extracted Deoxyribonucleic acid is too high, dilute it past using the TE buffer. Prepare a working stock of 30ng DNA for amplification.
Read more:
- What is a hot start PCR?
- Importance of Tris-EDTA (TE) buffer in DNA extraction
Determination:
Distension is not possible without the PCR DNA template. why are saying information technology as PCR Deoxyribonucleic acid template? because what DNA we are prepared (a purified 30ng DNA) can simply be used for the PCR amplification. For other genomic techniques, the concentration of the full DNA may vary.
[wp_ad_camp_2]
What Is A Template In Pcr,
Source: https://geneticeducation.co.in/what-are-the-properties-of-pcr-template-dna/
Posted by: martinthews1997.blogspot.com


0 Response to "What Is A Template In Pcr"
Post a Comment